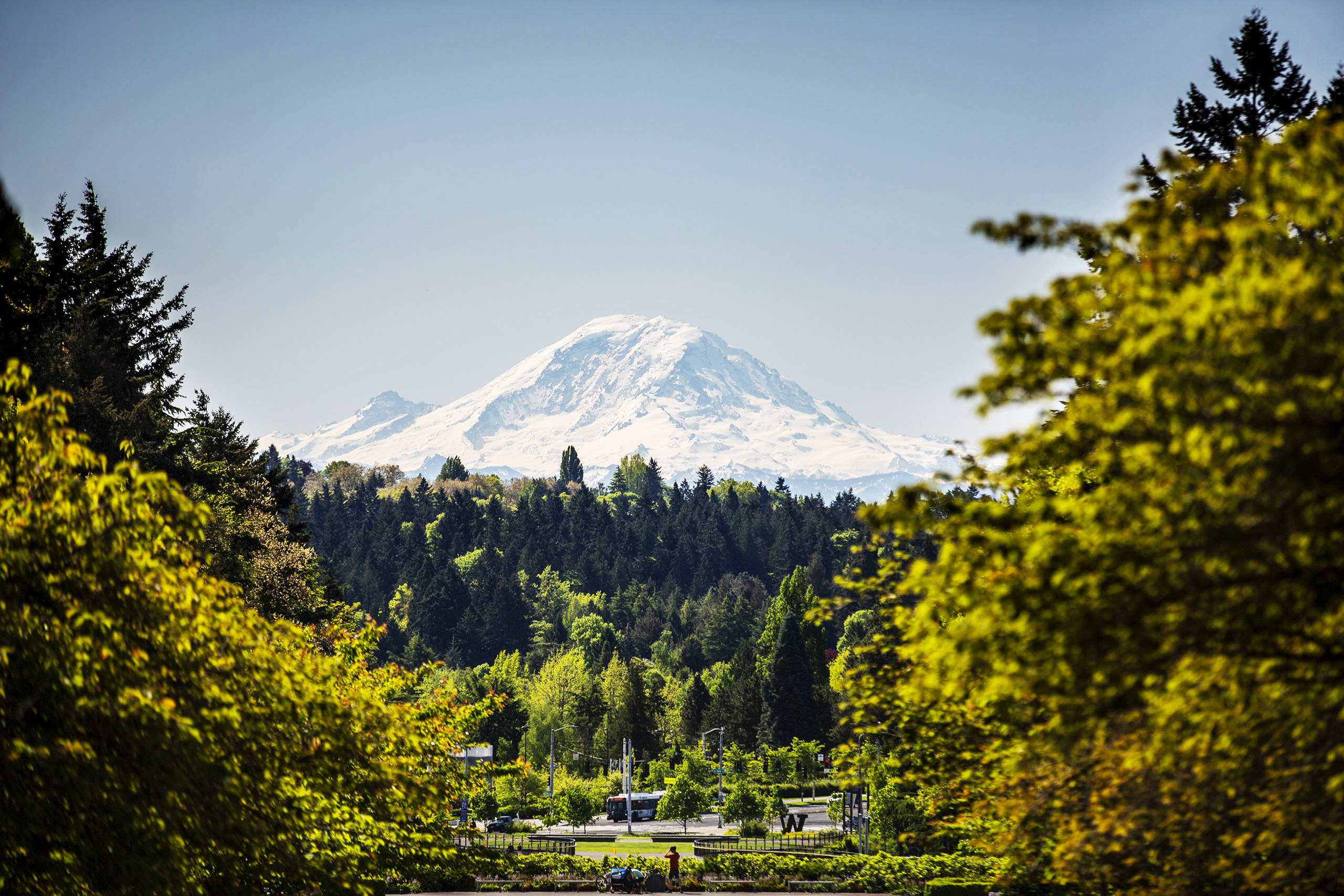
Derek
Stirewalt
M.D.

(206) 667-5386
Fred Hutch Cancer Center
1100 Fairview Ave. N, D5-112
Seattle, WA 98109
Photo: Fred Hutch
Education, Training, Board Certifications
- M.D., University of North Carolina at Chapel Hill
- Residency, University of North Carolina
- Fellowship in Hematology-Oncology, UW
- Medical Oncology, American Board of Internal Medicine
- Internal Medicine, American Board of Internal Medicine
Clinical Expertise
- Hematopoietic stem cell transplantation
- Hematologic malignancies
- Leukemia
Affiliations
- Fred Hutch Cancer Center - Faculty & Lab
- University of Washington
- Fred Hutch Cancer Center - Provider
Publications
Research and/or clinical interests
Dr. Derek Stirewalt studies the basic changes that occur in our blood stem cells as we age and their role in leukemia and other diseases. He uses cutting-edge technology to examine DNA, RNA and wide swaths of proteins to find new molecular calling cards as these diseases develop. Better understanding of the biological breakdown that leads to these diseases could lead to new tests for early detection and new therapies to halt or reverse them.
The overall goals of Dr. Stirewalt’s research are to identify novel biomarkers for hematopoietic malignancies and to understand how these biomarkers may function to promote the development of diseases. The purpose of this research is to improve the care of patients who are at risk for hematologic malignancies or who harbor these diseases.
Current projects include:
- Understand the biology driving clinical responses for patients with FLT3 and NPM1 mutations
Dr. Stirewalt’s lab was one of the first to examine the clinical significance of FLT3 mutations in AML. This work led to the recognition of FLT3 internal tandem duplications and NPM1 mutations as two of the most clinically utilized prognostic factors for AML patients, especially those with normal cytogenetics. Subsequent studies from the lab have found that the biology and clinical significance of these mutations differ depending on specific characteristics of the mutation (e.g., mutation size, location within the gene, allelic ratio), other cooperative molecular events (e.g., DNMT3A mutations), and age of the patient. Currently, his research projects investigating FLT3 and NPM1 focus on understanding the mechanisms driving the heterogeneous molecular biology and clinical significance of these mutations in AML. These studies are utilizing innovative technologies such as next generation sequencing, proteomics, and transfection/transduction models. - Identify novel AML prognostic biomarkers and therapeutic targets
Many AML patients do not harbor either NPM1 or FLT3 mutations; therefore, there is a need for additional prognostic biomarkers to better risk-stratify these patients. To identify novel prognostic factors and potential therapeutic targets, Dr. Stirewalt’s lab is examining differences in the global genomic and expression profiles between normal hematopoietic cells and AML blasts. Their previous studies have identified 13 genes with AML-specific expression changes, some of which may be novel prognostic factors and/or potential therapeutic targets, with several showing potential as novel adoptive immune targets. They are collaborating with other investigators to provide a more global assessment of the entire molecular landscape including protein expression and phosphorylation. - Understand the role of aging in the development of hematopoietic malignancies
As with most malignancies, the incidence of hematopoietic malignancies (AML, CML, MDS) increases with age, and therefore a better understanding of the molecular biology of the aging hematopoietic system may provide insight into hematopoietic transformation. Dr. Stirewalt’s lab identified a number of genes with age-related expression changes in normal hematopoietic progenitor/stem cells and AML blasts. Moreover, they and others have found that some of these age-related genes play critical roles in normal hematopoiesis and malignant transformation. Given the results from their previous work, they are expanding studies of aging to include a larger cohort of normal and malignant samples and to examine an even broader spectrum of expression changes (e.g., proteomics, alternative splicing, microRNA).